
  | THUNBERG, J., BERNARD, F., & GONCALVES, J. (June 2017). Distributed methods for synchronization of orthogonal matrices over graphs. Automatica, 80, 243–252. doi:10.1016/j.automatica.2017.02.025  Peer Reviewed verified by ORBi Peer Reviewed verified by ORBi |
  | BERNARD, F., SALAMANCA MINO, L., THUNBERG, J., Tack, A., Jentsch, D., Lamecker, H., Zachow, S., HERTEL, F., GONCALVES, J., & Gemmar, P. (May 2017). Shape-aware surface reconstruction from sparse 3D point-clouds. Medical Image Analysis, 38, 77-89. doi:10.1016/j.media.2017.02.005  Peer Reviewed verified by ORBi Peer Reviewed verified by ORBi |
  | BERNARD, F., Schmidt, F. R., THUNBERG, J., & Cremers, D. (2017). A Combinatorial Solution to Non-Rigid 3D Shape-to-Image Matching. In The proceedings of the IEEE Conference on Computer Vision and Pattern Recognition (CVPR). doi:10.1109/CVPR.2017.157  Peer reviewed Peer reviewed |
  | BERNARD, F. (2016). Novel Methods for Multi-Shape Analysis [Doctoral thesis, Unilu - University of Luxembourg]. ORBilu-University of Luxembourg. https://orbilu.uni.lu/handle/10993/29048 |
  | BERNARD, F., VLASSIS, N., Gemmar, P., HUSCH, A., THUNBERG, J., GONCALVES, J., & HERTEL, F. (2016). Fast Correspondences for Statistical Shape Models of Brain Structures. In SPIE Medical Imaging. doi:10.1117/12.2206024  Peer reviewed Peer reviewed |
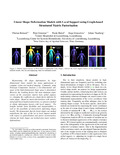  | BERNARD, F., Gemmar, P., HERTEL, F., GONCALVES, J., & THUNBERG, J. (2016). Linear Shape Deformation Models with Local Support using Graph-based Structured Matrix Factorisation. In Linear Shape Deformation Models with Local Support using Graph-based Structured Matrix Factorisation. doi:10.1109/CVPR.2016.607  Peer reviewed Peer reviewed |
  | SALAMANCA MINO, L., VLASSIS, N., DIEDERICH, N., BERNARD, F., & SKUPIN, A. (2015). Improved Parkinson’s disease classification from diffusion MRI data by Fisher vector descriptors. In Improved Parkinson’s disease classification from diffusion MRI data by Fisher vector descriptors (pp. 119-126).  Peer reviewed Peer reviewed |
  | BERNARD, F., SALAMANCA MINO, L., THUNBERG, J., HERTEL, F., GONCALVES, J., & Gemmar, P. (2015). Shape-aware 3D Interpolation using Statistical Shape Models. In Shape Symposium (pp. 11).  Peer reviewed Peer reviewed |
  | BERNARD, F., THUNBERG, J., Gemmar, P., HERTEL, F., HUSCH, A., & GONCALVES, J. (2015). A solution for Multi-Alignment by Transformation Synchronisation. In The proceedings of the IEEE Conference on Computer Vision and Pattern Recognition (CVPR). IEEE.  Peer reviewed Peer reviewed |
  | BERNARD, F., THUNBERG, J., SALAMANCA MINO, L., Gemmar, P., HERTEL, F., GONCALVES, J., & HUSCH, A. (2015). Transitively Consistent and Unbiased Multi-Image Registration Using Numerically Stable Transformation Synchronisation. MIDAS Journal.  Peer reviewed Peer reviewed |
  | HUSCH, A., Gemmar, P., Lohscheller, J., BERNARD, F., & HERTEL, F. (2015). Assessment of Electrode Displacement and Deformation with Respect to Pre-Operative Planning in Deep Brain Stimulation. In H. Handels, T. M. Deserno, H.-P. Meinzer, ... T. Tolxdorff (Eds.), Bildverarbeitung für die Medizin 2015 (pp. 77-82). Springer Berlin Heidelberg. doi:10.1007/978-3-662-46224-9_15  Peer reviewed Peer reviewed |
  | HERTEL, F., HUSCH, A., Dooms, G., BERNARD, F., & Gemmar, P. (2015). Susceptibility-Weighted MRI for Deep Brain Stimulation: Potentials in Trajectory Planning. Stereotactic and Functional Neurosurgery, 93 (5), 303-308. doi:10.1159/000433445  Peer Reviewed verified by ORBi Peer Reviewed verified by ORBi |
  | BERNARD, F., Gemmar, P., HUSCH, A., Saleh, C., Neb, H., Dooms, G., & HERTEL, F. (2014). Improving the Consistency of Manual Deep Brain Structure Segmentations by Combining Variational Interpolation, Simultaneous Multi-Modality Visualisation and Histogram Equilisation. Biomedizinische Technik. Biomedical Engineering, 59 (1), 131-134. doi:10.1515/bmt-2014-5008  Peer reviewed Peer reviewed |
  | BERNARD, F., Gemmar, P., HUSCH, A., & HERTEL, F. (2014). An Extensible Development Environment for 3D Segmentations based on Active Shape Models. In Shape Symposium (pp. 39).  Peer reviewed Peer reviewed |